Unified AI-Powered Platform for Computational Biology
Neurosnap is a web-based bioinformatics platform that delivers advanced computational biology tools through a single, intuitive AI-driven web interface. From protein folding and molecular docking to RNA-seq analysis and in silico mutagenesis, Neurosnap removes the complexity of installing, maintaining, and scaling local bioinformatics software.
Designed for both academic and professional researchers, Neurosnap enables scientists to focus on discovery, not infrastructure.

Why Neurosnap?
Modern bioinformatics workflows often require high-performance computing, complex software installations, and deep programming expertise. Neurosnap changes this by providing a zero-code (or low-code) environment where advanced analyses can be executed directly in the browser.
Explore 100+ curated, validated bioinformatics tools.
Gain access to a trusted collection of 100+ published, state-of-the-art models and workflows with proven performance records.
Work entirely in your browser—no setup, no installs.
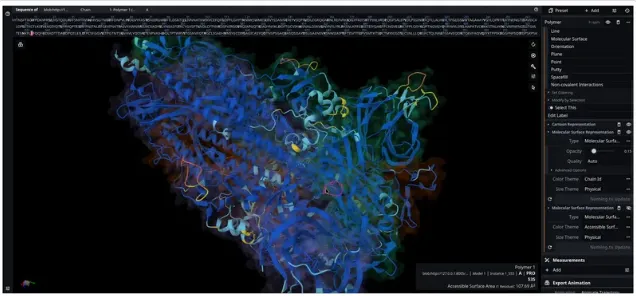
Perform folding, docking, and simulations seamlessly in our web workspace, analyze your results, fine-tune them easily, and share updates with your team instantly.
Design intelligent pipelines built around your workflow.
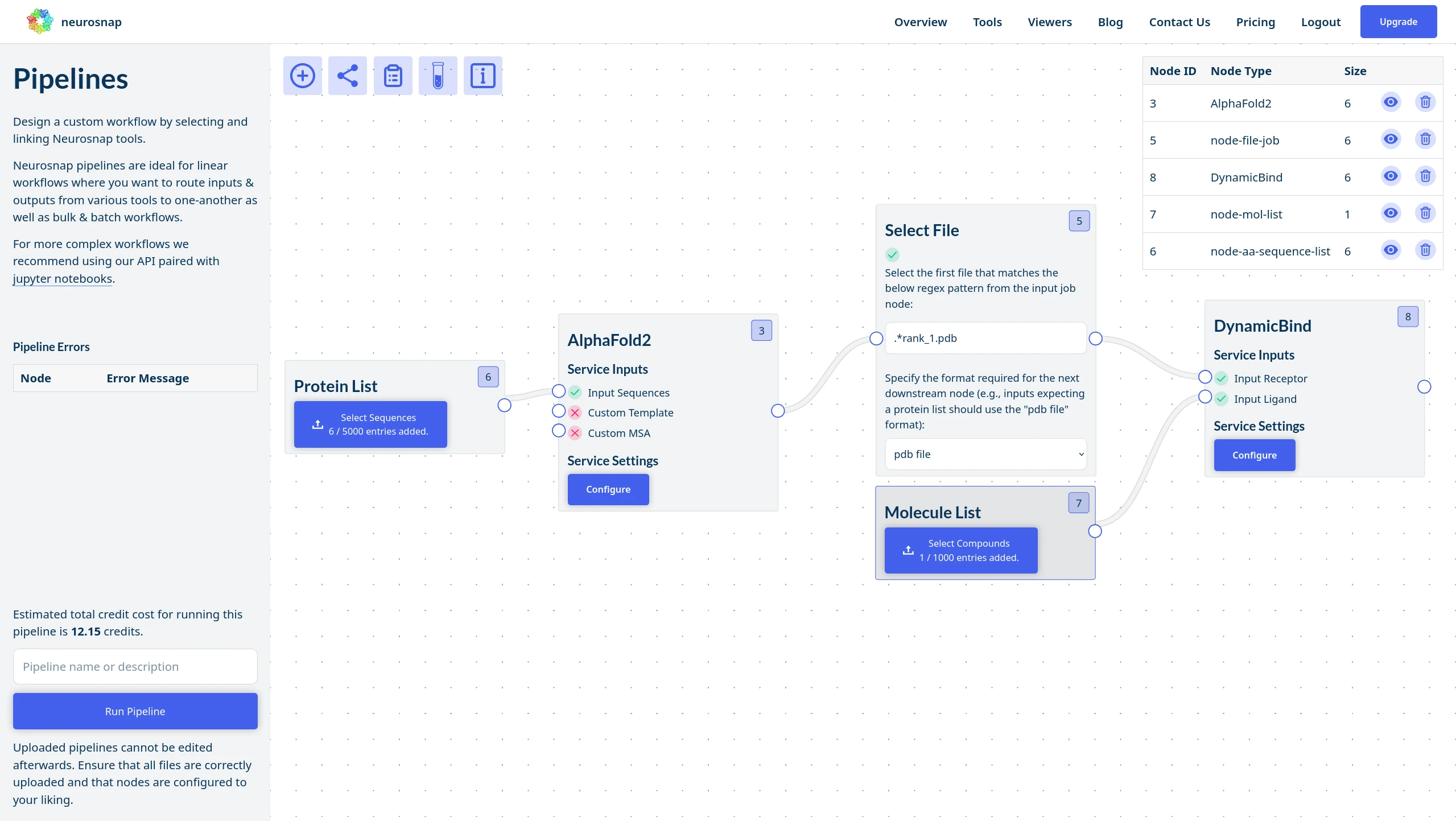
NeuroSnap Pipelines allow you to seamlessly connect tools, route outputs downstream, and automate decisions using custom logic.
Let the API handle the heavy lifting.
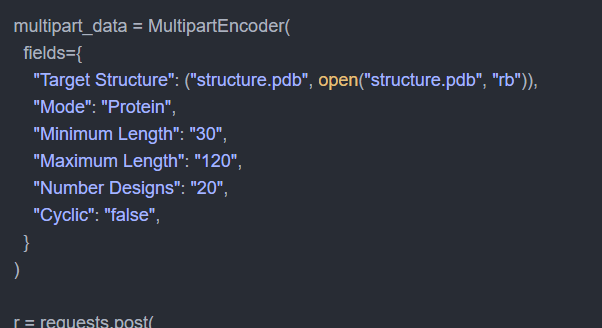
Programmatically submit jobs, stream results, and integrate NeuroSnap into your existing pipelines. API examples are available for all tools.
With Neurosnap, researchers can:
Discover Powerful Features of Neurosnap for Smarter Scientific Decisions
NeuroSnap enables researchers to execute advanced computational biology workflows through an integrated, AI-powered platform. Each capability combines validated scientific methods with modern machine-learning models to accelerate discovery while maintaining analytical rigor.
How NeuroSnap Works?
NeuroSnap simplifies complex computational biology workflows by combining AI bioinformatics software, cloud infrastructure, and automated pipelines into a single, zero-code environment.
What Can You Do with NeuroSnap
Advanced Bioinformatics with Neurosnap AI ?
AI Models & Engines Powering NeuroSnap
Antibody Design
AI-powered antibody engineering workflows
NeuroSnap enables antibody discovery and optimization by modeling antigen–antibody interactions and multi-chain assemblies. These workflows support rational antibody design with a focus on binding specificity and developability.
Best used for:
- CDR loop modeling and refinement
- Antigen-specific antibody design
- Affinity maturation and interaction analysis
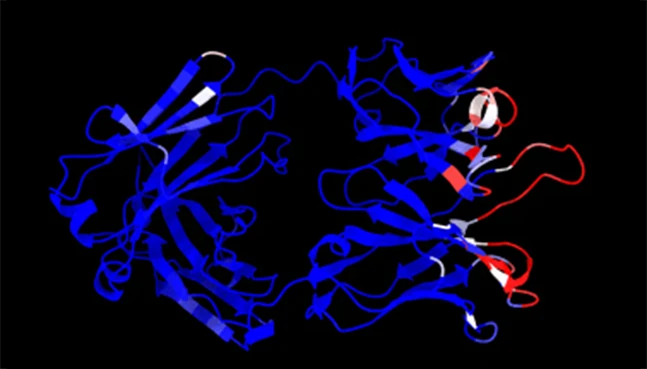
Protein Annotation
Automated functional and domain-level interpretation
NeuroSnap annotates protein sequences and structures with functional domains, motifs, and biological context, enabling rapid interpretation without manual curation.
Best used for:
- Domain and motif identification
- Structure–function insights
- Pathway and ontology mapping
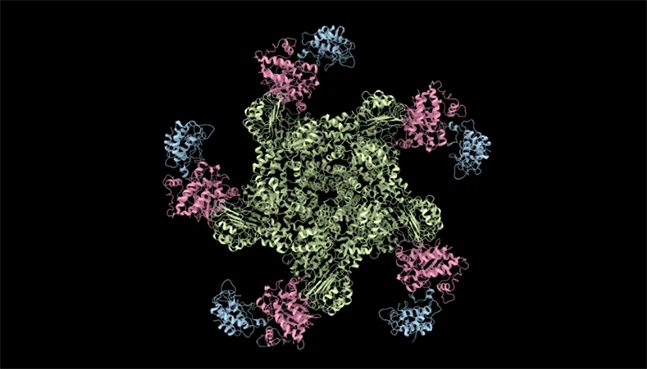
Protein Clustering
Large-scale protein similarity analysis
NeuroSnap groups proteins based on sequence and structural similarity, enabling efficient dataset organization and comparative proteomics at scale.
Best used for:
- Protein family classification
- Redundancy reduction
- Comparative protein analysis
Protein Localization
Subcellular targeting prediction
NeuroSnap predicts the most likely cellular location of proteins, providing biological context that informs experimental design and downstream validation.
Best used for:
- Subcellular localization analysis
- Functional role interpretation
- Experimental planning
Signal Peptide Detection
Secretory pathway and targeting analysis
NeuroSnap identifies signal peptides and cleavage sites to determine whether proteins are secreted or membrane-associated.
Best used for:
- Secreted protein discovery
- Membrane protein screening
- Expression system selection
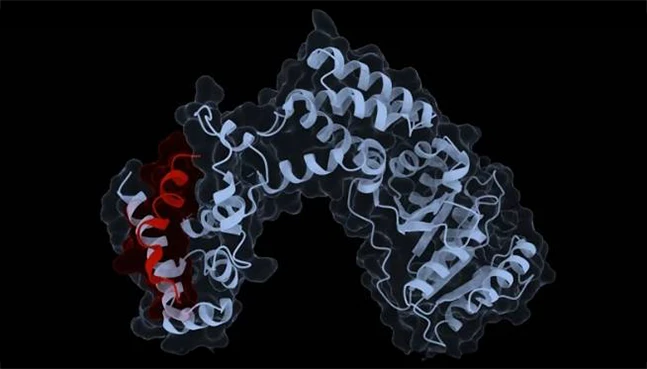
Protein Solubility
Developability-focused solubility assessment
NeuroSnap evaluates solubility and aggregation risk to help prioritize protein variants with higher experimental success rates.
Best used for:
- Expression feasibility screening
- Aggregation risk reduction
- Variant prioritization

Protein Expression
Host-aware expression optimization
NeuroSnap analyzes sequence features and codon usage to predict protein expression efficiency across common host systems.
Best used for:
- Host-specific expression planning
- Codon optimization insights
- Production success prediction
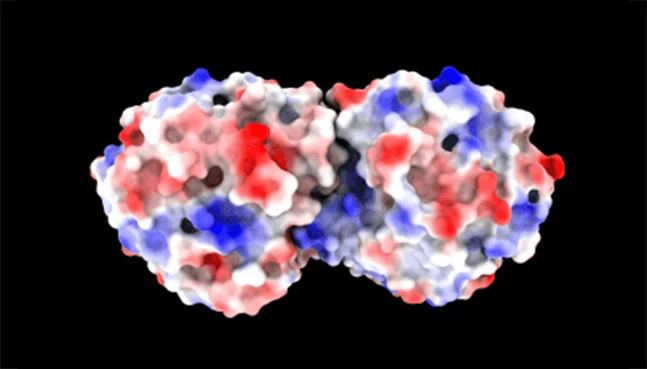
Toxicity Prediction
Early-stage safety assessment
NeuroSnap screens proteins and peptides for potential toxicity and off-target effects, enabling safer design decisions earlier in the pipeline.
Best used for:
- Therapeutic candidate screening
- Risk mitigation
- Lead prioritization
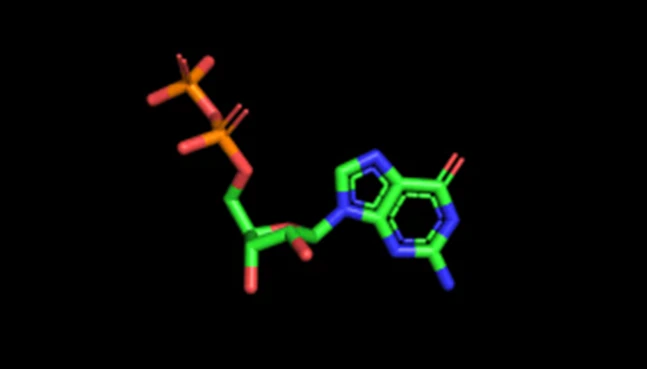
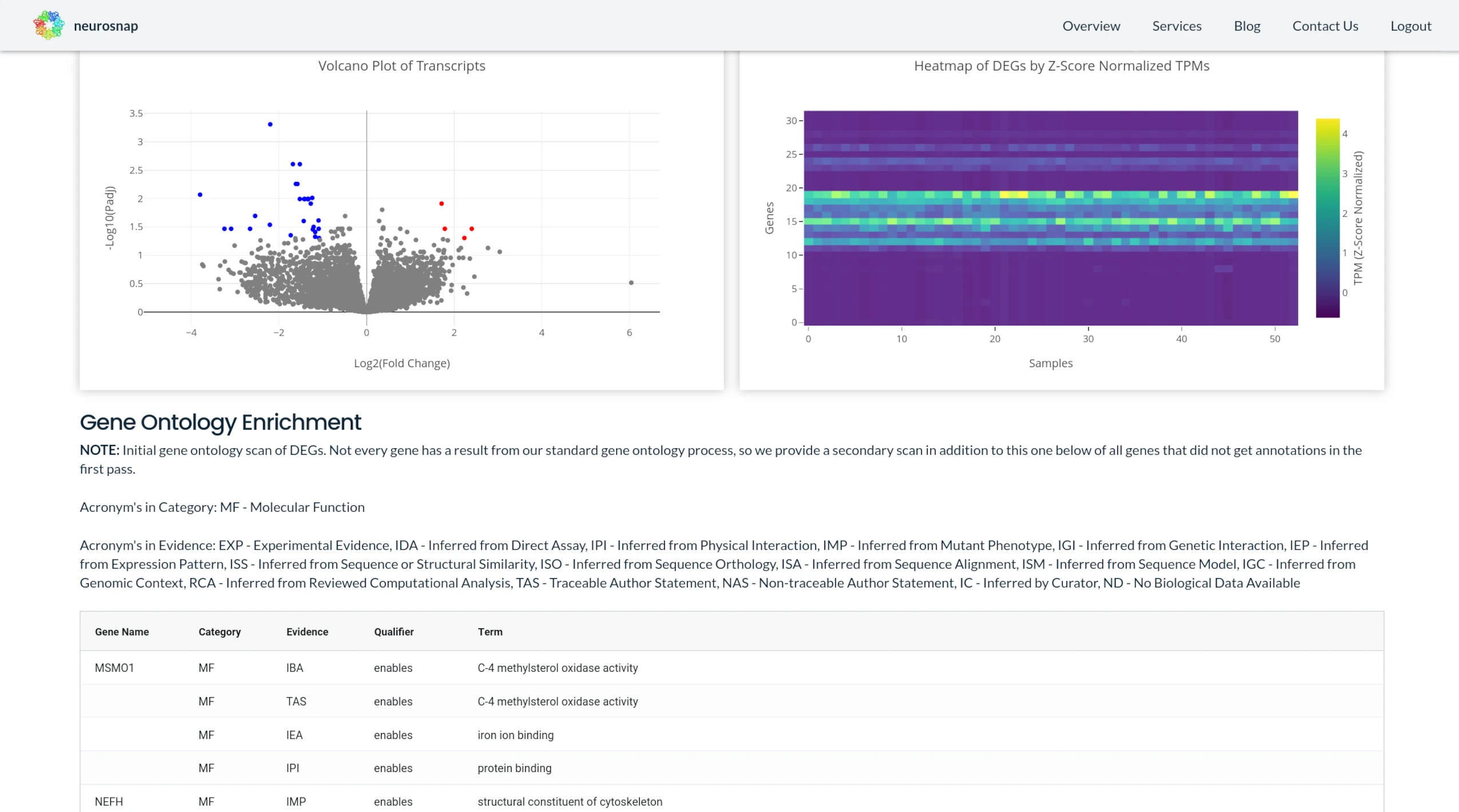
RNASeq
End-to-end RNA sequencing analysis
NeuroSnap processes raw RNASeq data into actionable gene expression insights using automated pipelines and AI-assisted interpretation.
Best used for:
- Gene expression profiling
- Differential expression analysis
- Transcript discovery
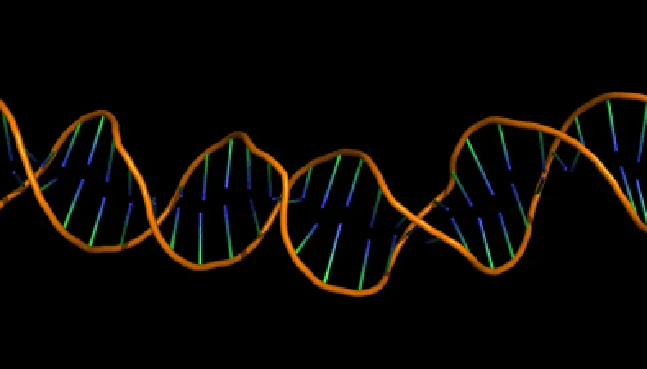
Transcriptome Analysis
Systems-level gene regulation insights
NeuroSnap integrates RNASeq outputs to deliver a comprehensive view of transcript-level regulation, alternative splicing, and biological response patterns.
Best used for:
- Pathway-level expression analysis
- Condition-based transcriptome comparison
- Biomarker discovery
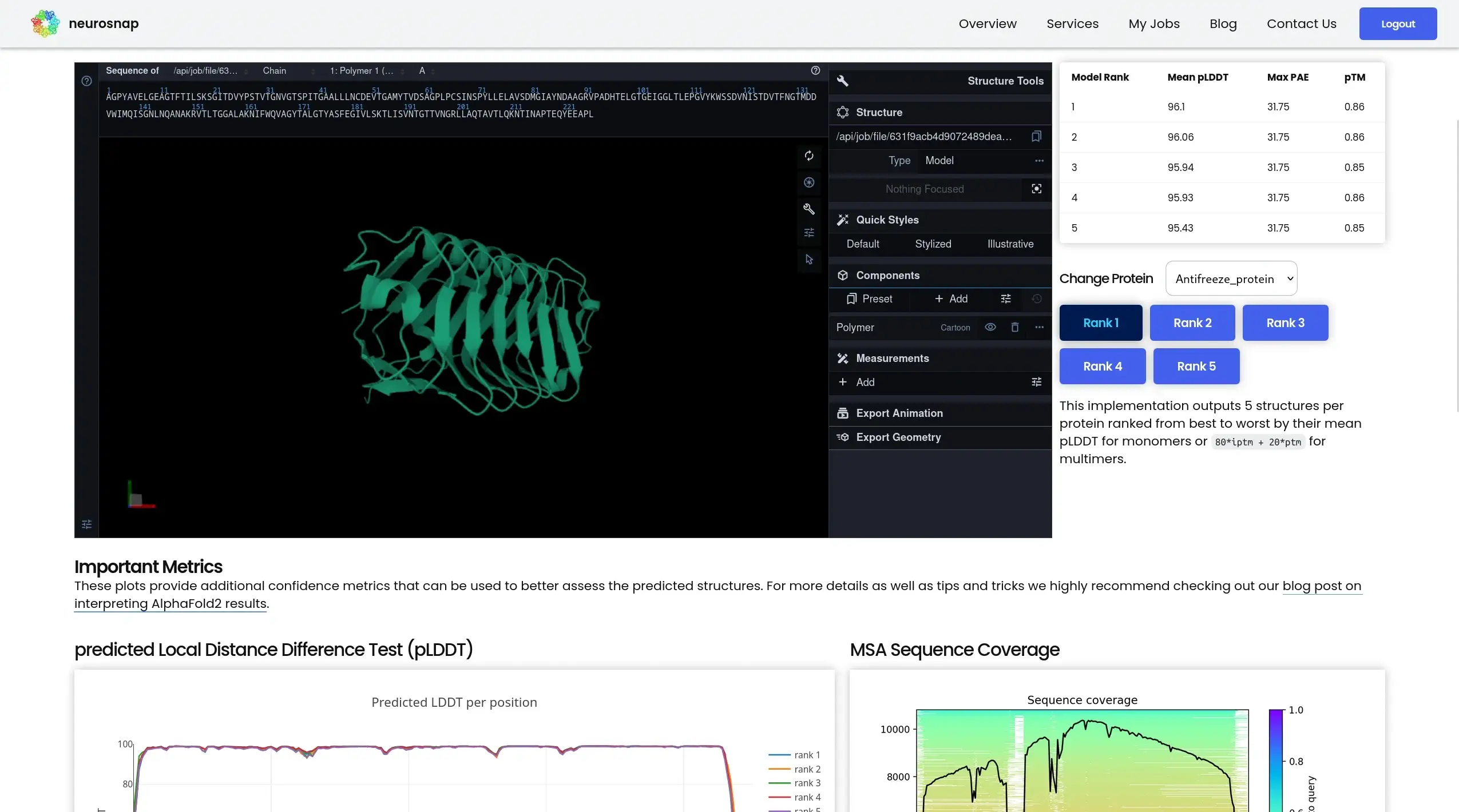
AlphaFold2
High-accuracy protein structure prediction
AlphaFold2 predicts three-dimensional protein structures from amino acid sequences using deep learning trained on known structural data. It is widely trusted for high-confidence structure generation and serves as a foundational tool for structural biology and downstream computational analyses.
Best used for:
- Benchmark-quality protein structures
- Structural interpretation and validation
- Input for docking and molecular simulations
ESMFold
Fast structure prediction using protein language models
ESMFold leverages large protein language models to quickly predict protein structures. It is optimized for speed and scalability, making it ideal for large datasets and rapid exploratory studies.
Best used for:
- High-throughput structure prediction
- Rapid prototyping and screening
- Preliminary structural analysis
Boltz-1
Next-generation structure prediction for complex systems
Boltz-1 is designed to predict structures of complex biological assemblies, including protein complexes and challenging targets. It offers enhanced modeling capabilities for multi-chain and interaction-heavy systems.
Best used for:
- Protein complexes
- Interaction-focused studies
- Advanced structural modeling
AlphaFlow
Next-generation structure prediction for complex systems
AlphaFlow applies flow-matching principles to structure prediction, allowing efficient sampling of protein conformations. It complements traditional folding tools by offering alternative structural perspectives.
Best used for:
- Structural diversity exploration
- Conformation sampling
- Comparative structure analysis
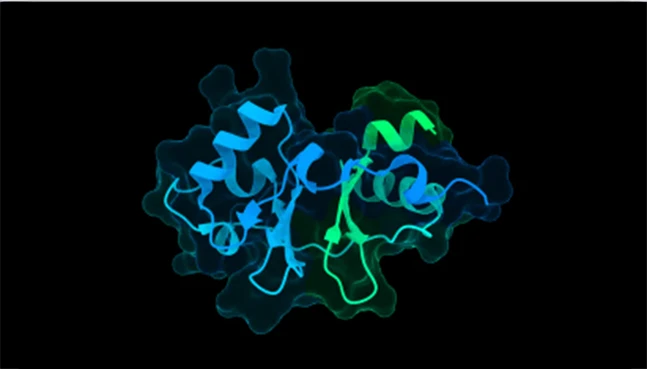
Chai-1
AI-assisted multi-chain and interaction modeling
Chai-1 focuses on modeling biomolecular interactions and assemblies, addressing challenges in predicting multi-chain and interdependent protein systems.
Best used for:
- Multi-chain protein systems
- Interaction modeling
- Complex biological assemblies
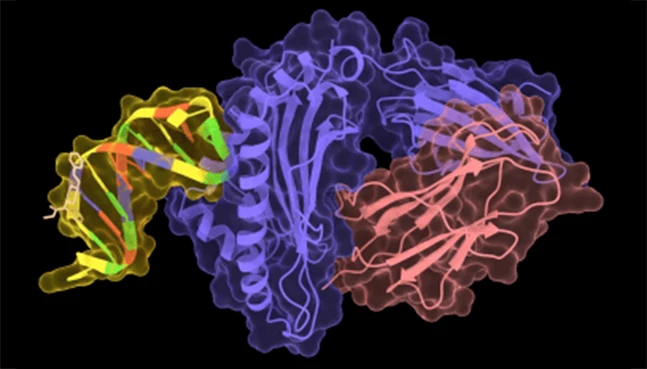
ProteinMPNN
Structure-guided protein sequence design
ProteinMPNN designs amino acid sequences that are compatible with a given protein backbone. It enables rational protein design by optimizing sequences while preserving structural integrity.
Best used for:
- Protein redesign
- Functional variant generation
- Backbone-constrained sequence optimization
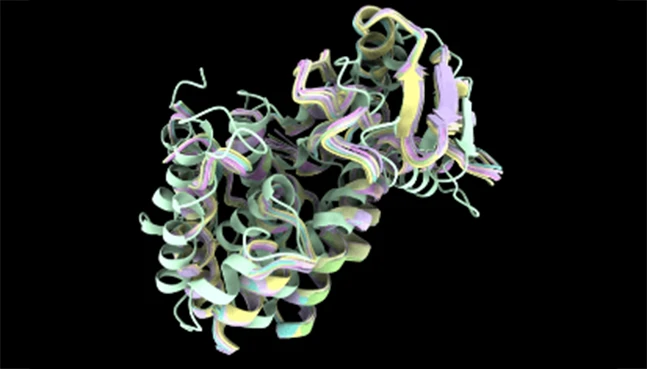
ThermoMPNN
Stability-focused protein optimization
ThermoMPNN extends sequence design by prioritizing thermodynamic stability and robustness. It is particularly useful for improving protein performance under demanding conditions.
Best used for:
- Thermostability optimization
- Industrial and enzyme applications
- Stability-driven variant screening
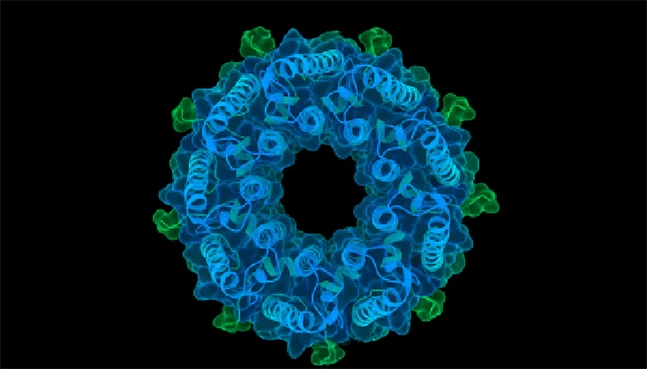
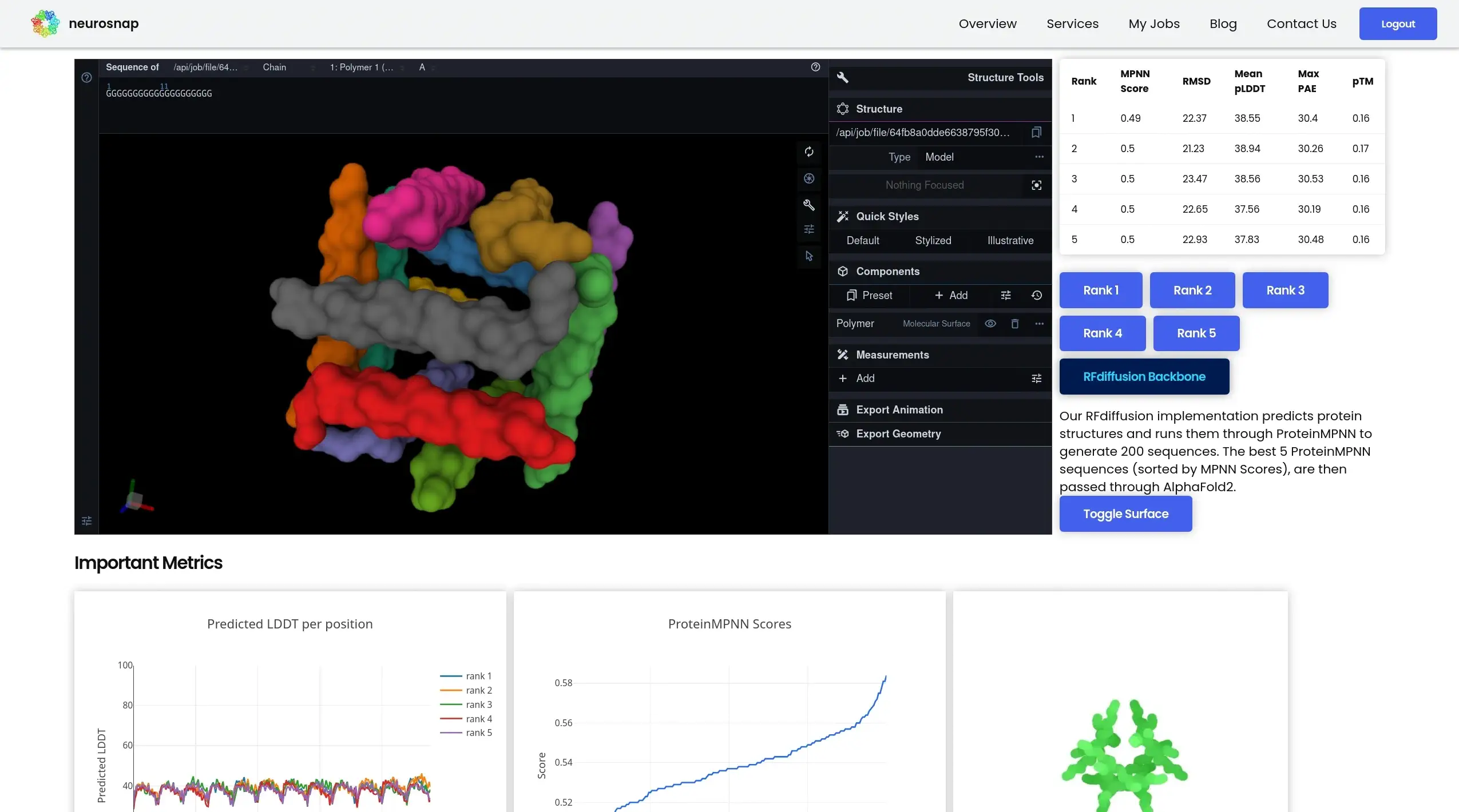
RFdiffusion
Generative protein structure design
RFdiffusion generates entirely new protein structures or scaffolds using diffusion-based generative modeling. It enables exploration beyond known protein families for innovative design applications.
Best used for:
- De novo protein design
- Scaffold generation
- Binder and motif creation
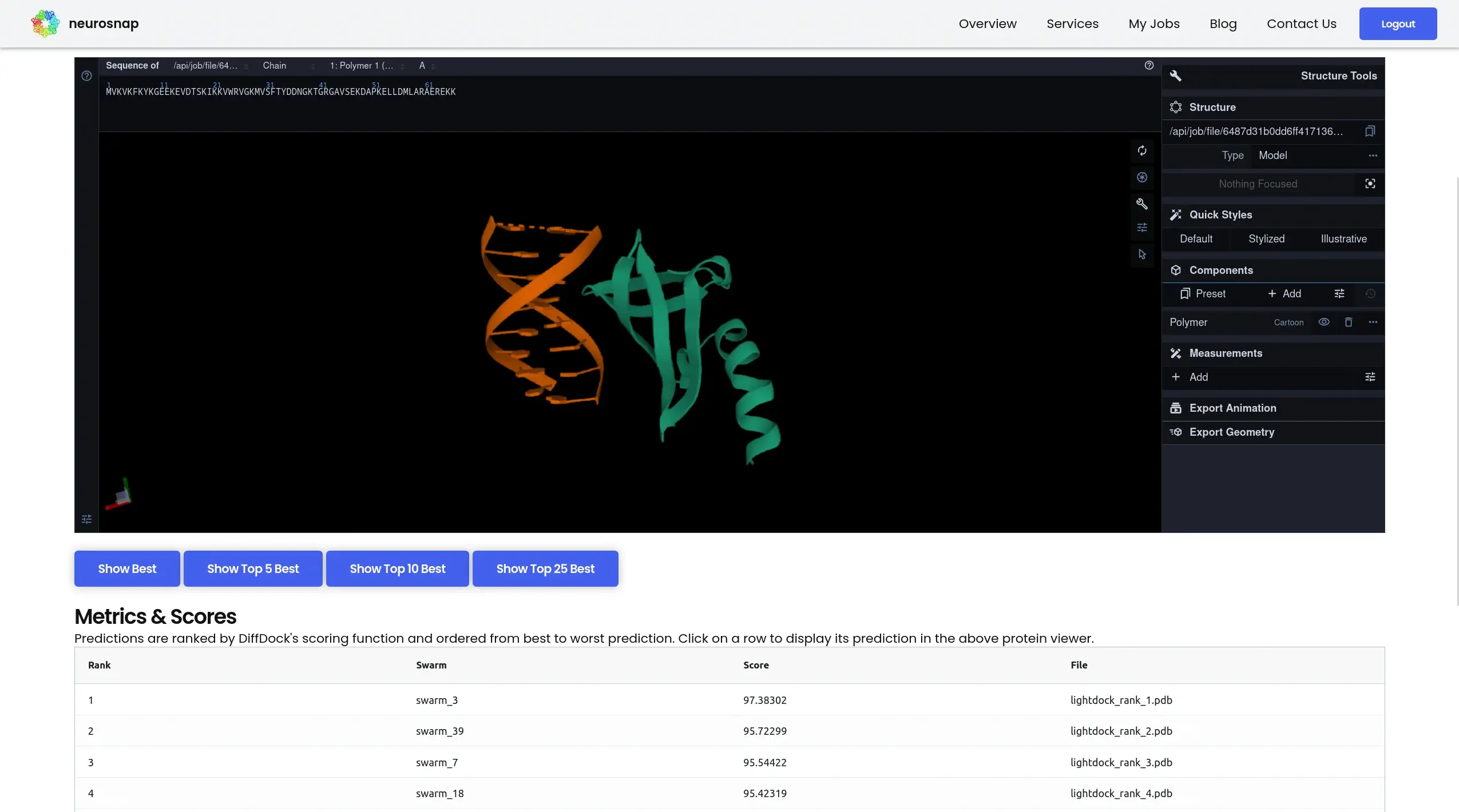
GNINA
Deep learning–enhanced molecular docking
GNINA integrates convolutional neural networks into molecular docking workflows to improve pose prediction and scoring accuracy.
Best used for:
- Protein–ligand docking
- Drug discovery and screening
- Binding affinity estimation
PDBFixer
Structure preparation and correction utility
PDBFixer cleans and repairs protein structures by fixing missing residues, atoms, and formatting issues—ensuring models are ready for docking, simulations, or further computational workflows.
Best used for:
- Structure preprocessing
- Model cleanup and validation
- Simulation-ready preparation

FAQ’s
Neurosnap bioinformatics tools are ideal for:
- Academic researchers and students
- Labs without dedicated HPC resources
- Protein and enzyme engineering teams
- Drug discovery and molecular design projects
- Synthetic biology and transcriptomics research
The platform supports:
- Protein structure prediction
- Protein engineering and variant analysis
- Binding and interaction studies
- Molecular docking
- Molecular dynamics simulations